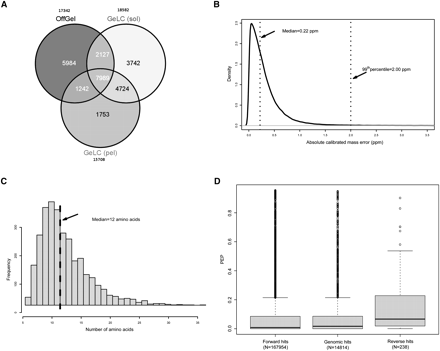
Overview of the proteomics results. (A) Application of complementary biochemical workflows for protein extraction and peptide separation led to enhanced proteome coverage. (sol) Soluble fraction; (pel) pellet. (B) Peptides were detected with a mean absolute mass deviation of 0.345 ppm. (C) Peptide sequences that mapped to the genome translation (“genomic peptides”) had a median size of 12 amino acids. (D) Distribution of posterior error probabilities (PEP) was markedly different in the genomic and reversed peptide sequences.











