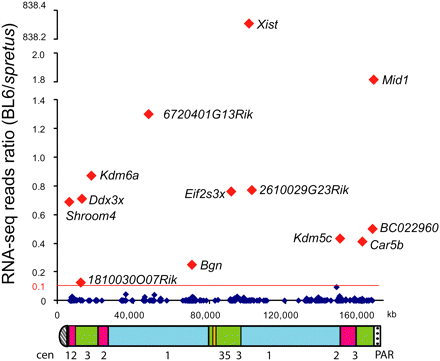
Distribution of escape genes on the mouse X chromosome. The ratio between RNA-seq reads from BL6 (inactive X) and M. spretus (active X) is graphed for each gene against its location (UCSC Genome Browser, Mouse July 2007 [mm9] assembly). (Red) Genes with an average expression level from the inactive X higher than 10% of the active X; (dark blue) other genes. Underneath, an ideogram of the mouse X chromosome indicates regions that correspond to evolutionary strata (1, 2, 3, and 5) defined in human (Ross et al. 2005). Genes in strata 4 are apparently deleted in mouse. (Cen) Centromere; (PAR) pseudoautosomal region.











