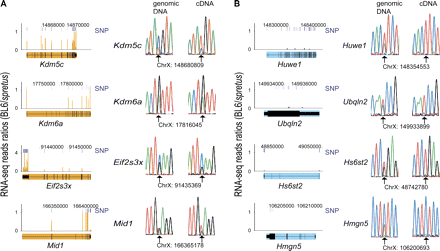
RNA-seq identification and conventional sequencing validation of the X-inactivation status of mouse genes. (A) Four genes previously known to escape X inactivation (Kdm5c, Kdm6a, Eif2s3x, Mid1; filled in orange) correctly identified by RNA-seq; (B) four examples of genes subject to X inactivation (Huwe1, Ubqln2, Hs6st2, Hmgn5; filled in light blue) identified by RNA-seq. (Left) RNA-seq data are graphed as the sequence read ratio (orange bars) between BL6 (inactive X) and M. spretus (active X) at each SNP (short, dark-blue bars). A few SNPs had ratios higher than those at the maximum scale (Supplemental Table S1). Data were uploaded into UCSC Genome Browser (Mouse July 2007 [mm9] assembly). (Top) Chromosome X coordinates. (Right) Validation of SNPs and of bi-allelic (A) or mono-allelic (B) gene expression by conventional sequencing of PCR and RT-PCR products. Chromatograms are shown for both genomic DNA and cDNA with SNP coordinate underneath. (Arrows) Heterozygous bases.











