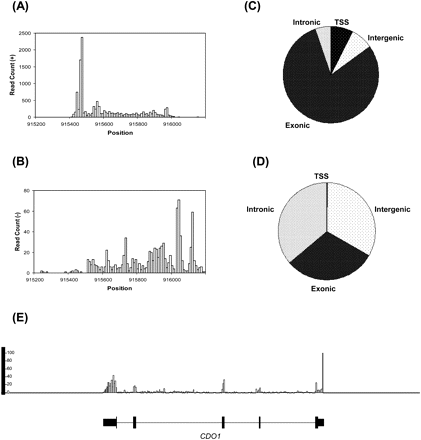
Coverage of reads aligned in the + direction (A) and − direction (B) binned in 10-nt intervals is exemplified across the HTA1 gene (location: chr4 915,521–915,919; transcription direction: +). (C) Assignment of uniquely aligned S. cerevisiae polyA+ RNA reads to the categories shown. TSS (transcription start site) category refers to regions 200 nt upstream of annotated coding region start sites; 7.4% of reads map to the yeast TSS regions. Reads in the S. cerevisiae intergenic regions include reads aligning mostly to the potentially transcriptionally active transposon repeats. (D) Assignment of uniquely aligned human liver polyA+ RNA reads to the categories shown. TSS category refers to regions 200 nt upstream and downstream of the annotated RefSeq transcription start sites; 0.2% of the reads map to the human TSS regions. (E) Human liver polyA+ RNA reads uniquely aligning to human genome was binned at 50-nt intervals and visualized using the Integrated Genome Browser. Panel exemplifies the distribution of reads across the human CDO1 (location: chr5 115,168,329–115,180,304; transcription direction: −) gene's exonic (thick bars) and intronic (thin lines) regions. Y-axis indicates the number of reads per bin.











