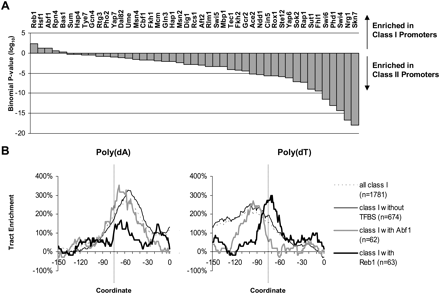
Poly(dA:dT) enrichments occur independently of transcription factor binding sites. (A) Ranking of transcription factors by overrepresentation of bound and functionally conserved sites in class I promoters based on annotated sites from MacIssac et al. (2006). P-values were computed by modeling the distribution of TFBS between class I and class II promoters using a binomial distribution. The vertical axis gives the logarithm of the cumulative binomial probably of having the observed number of TFBS in a given class. Overrepresentation in class I promoters is shown by positive values and in class II promoters by negative values. (B) Poly(dA:dT) tract enrichments (length ≥ 4) for class I promoters and subsets thereof: Abf1-containing promoters, Reb1-containing promoters, and TFBS-depleted promoters. TFBS-depleted promoters were selected by excluding promoters containing moderately bound (P < 0.005) binding sites (no conservation requirement) for any of 118 TFs.











