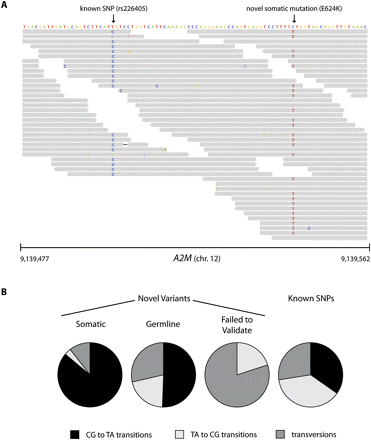
Validation of somatic mutations. (A) Sequence variants in exon 16 of A2M in melanoma short-term culture M970109. Illumina 51-mer reads are shown as gray boxes (arrowheads denote directionality of reads; nonreference bases are colored and shaded according to their quality scores). One variant (T → C) is homozygous and corresponds to a known SNP in dbSNP (Sherry et al. 2001). The other variant (C → T) is heterozygous and corresponds to a missense E624K mutation. This mutation was validated and confirmed to be somatic. (B) Distribution of transitions and transversions in sequence variants genotyped by Sequenom. Validated somatic mutations were largely CG to TA transitions (86%), representative of mutations induced by UV damage (Drobetsky et al. 1987). In contrast, only 53% of novel germline variants and 0% of variants that failed to validate were CG to TA transitions. Thirty six percent of all known SNPs detected by Illumina in the 10 melanoma samples were CG to TA transitions with respect to human genome reference hg18.











