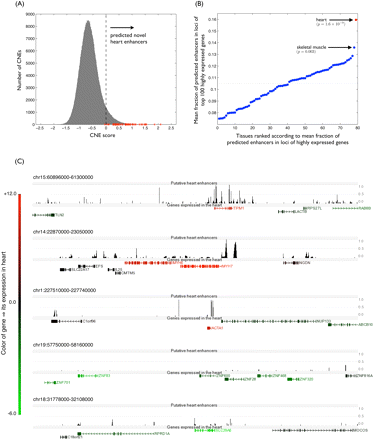
(A) Distribution of heart scores of CNEs. Scores assigned by the classifier for all tested CNEs are shown here. We use zero as a cutoff (Methods) for putative enhancers (dotted line). (Red) Scores of the training enhancer set. (B) Mean fraction of high-scoring CNEs in loci of genes highly expressed in each tissue. Tissues are sorted based on the mean fraction of putative heart enhancers in their loci. P-values were computed using a rank sum test; heart tissue had the most significant P-value of 1.6 × 10−9. (C) (Black peaks) Snapshots of genome-wide view of predictions near genes. The score returned by the classifier is transformed to lie between 0 and 1, with numbers >0.5 indicating the occurrence of a putative heart enhancer. The color and shade of the gene transcript depict the type and level of gene expression, respectively: (red) genes highly expressed in the heart; (green) repressed genes. Genes highly expressed in the heart have typically more enhancers in their loci (top three genomic regions), while genes repressed or not expressed in the heart have fewer predictions in their loci. (All elements in the training set are excluded in these figures.)











