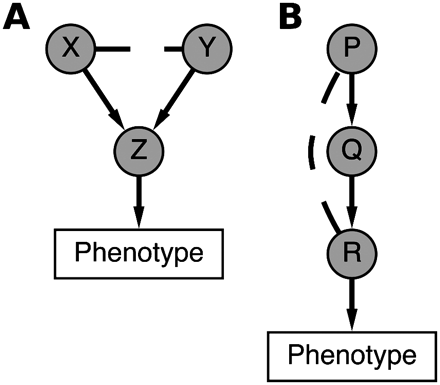
Model for between-pathway and within-pathway genetic interactions for high-dimensional morphological data. (A) A between-pathway genetic interaction for genes X and Y is said to occur when single-knockdown of either gene does not result in a mutant phenotype, but the double-knockdown X/Y does. For this study, X and Y are RhoGAPs, the gene Z is a RhoGTPase, and the mutant phenotype is morphological similarity at the single-cell level to overexpression of Z. In this way, identification of between-pathway genetic interactions in our combinatorial knockdown data set corresponds to prediction of RhoGAP/RhoGTPase-signaling interactions (see text). (B) A within-pathway genetic interaction between genes P and R is said to occur when the double-knockdown P/R bears significant morphological similarity to either the single-knockdown for P or R, but not both. In our study, P, Q, and R are RhoGAPs, and identification of within-pathway genetic interactions corresponds to complex hierarchical relations between RhoGAPs.











