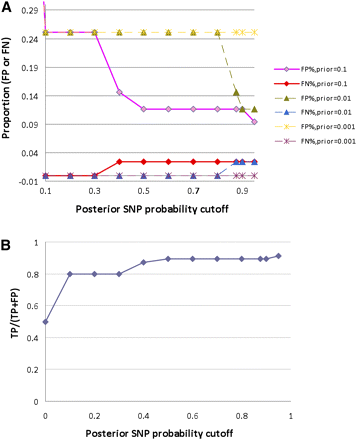
The validation results in S. aureus data with at least three variant reads when three different sets of priors were used for tuning purposes. We used three sets of priors (Supplemental Table S2) in Equation 5 for SNP probability assessment. (A) The false-positive rate (FP) and false-negative rate (FN) can be evaluated using our defined SNPs and errors (described in Methods). The results indicate that a 10% false-positive rate and a 5% false-negative rate can be achieved when using either the “set 1” or the “set 2” parameters, while “set 1” enables a smoother resolution. (B) The FP/[FP + true-positives (TP)] is plotted against the posterior SNP probability cutoff for results obtained using “set 1” priors.











