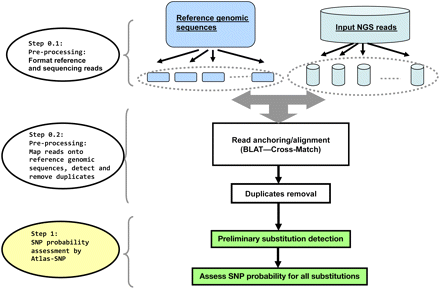
The overall workflow of the Atlas-SNP2 package. The reference genomic sequence and reads undergo an initial data processing step, whereby the reference sequence is split into smaller pieces and the reads into smaller batches. A combined BLAT and Cross_Match analysis was used to anchor and align reads back to the reference positions. All of the single nucleotide mismatches are parsed and assessed for their probabilities of being SNPs using the Atlas-SNP2 core statistical methods.











