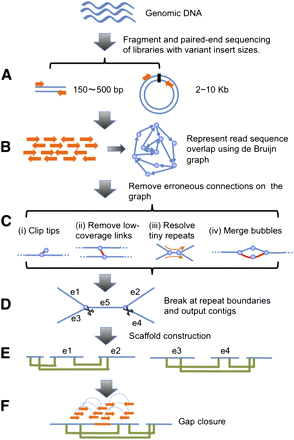
Schematic overview of the assembly algorithm. (A) Genomic DNA was fragmented randomly and sequenced using paired-end technology. Short clones with sizes between 150 and 500 bp were amplified and sequenced directly; while long range (2–10 kb) paired-end libraries were constructed by circularizing DNA, fragmentation, and then purifying fragments with sizes in the range of 400–600 bp for cluster formation. (B) The raw or precorrected reads were then loaded into computer memory and de Bruijn graph data structure was used to represent the overlap among the reads. (C) The graph was simplified by removing erroneous connections (in red color on the graph) and solving tiny repeats by read path: (i) Clipping the short tips, (ii) removing low-coverage links, (iii) solving tiny repeats by read path, and (iv) merging the bubbles that were caused by repeats or heterozygotes of diploid chromosomes. (D) On the simplified graph, we broke the connections at repeat boundaries and output the unambiguous sequence fragments as contigs. (E) We realigned the reads onto the contigs and used the paired-end information to join the unique contigs into scaffolds. (F) Finally, we filled in the intrascaffold gaps, which were most likely comprised by repeats, using the paired-end extracted reads.











