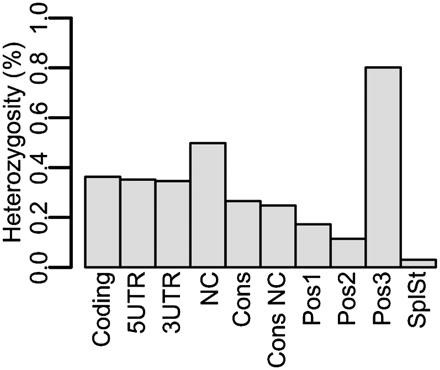
Our fly was compared with the dm3 reference across various annotated regions using SSAHA-SNP. Within annotated regions the variation rate matches previous Drosophila variation findings. The most conserved regions were the first (Pos1) and second (Pos2) codon positions, conserved sequence (Cons), and splice-site junctions (SplSt). The most variable region was the third codon position (Pos3). For comparison, annotated protein-coding regions (Coding), 5′ untranslated regions (5UTR), 3′ untranslated regions (3UTR), noncoding sequence (NC), and conserved noncoding sequence (Cons NC) were included.











