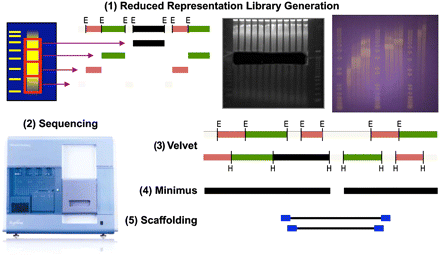
A schematic overview of the sequencing and assembly methods used to generate our de novo fly assembly. (1) Reduced representation libraries were created by digestion of genomic DNA with two restriction enzymes separately. Shown are a single library's gel slice and a subsequent second purification step to ensure library fragment fidelity. Resolution on an agarose gel allowed for libraries to be selected between 1–4 kb, 4–7 kb, 7–9 kb, and 9–30 kb in size for each enzyme independently. (2) Each library was then sequenced independently on the Illumina Genome Analyzer. (3) The short-read libraries were then assembled using Velvet. (4) Overlapping contigs from all eight libraries were merged using the lightweight assembler Minimus. (5) Finally, genomic paired-end short sequence reads were incorporated into the assembly process to order and orient the contigs generated in previous steps.











