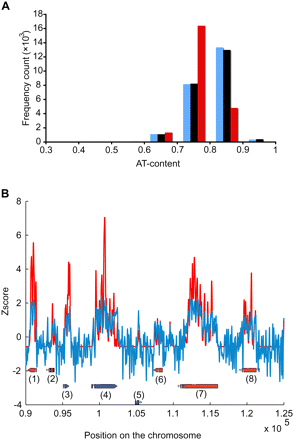
P. falciparum genome and its effect on nucleosome positioning. (A) AT-content distributions across the genome, FAIRE-enriched and MAINE-enriched regions were compared. AT-content was calculated in a 1000-bp-long sliding window for the P. falciparum genome (blue) and regions covered by FAIRE- and MAINE-seq (black and red, respectively). AT-content is similar genome-wide and in FAIRE-covered regions. On the contrary, regions enriched by MAINE are influenced toward less AT-rich regions, which is in agreement with the low affinity of nucleosomes for AT-rich sequences. Same distributions were obtained using sliding windows of various sizes (10, 50, 100, 1000, and 20,000 bp) (data not shown) further validating the efficiency of Illumina technology for the P. falciparum AT-rich genome. (B) Correlation between experimental and in silico nucleosome positioning was explored. MAINE profile (red) and DNA bending free energy profile (blue) were z-normalized and overlaid for the region [90,000–125,000] of chromosome 1. Oriented boxes represent gene models retrieved from http://www.plasmodb.org: (1) PFA0095c; (2) PFA0100c; (3) PFA0105w; (4) PFA0110w; (5) PFA0115w; (6) PFA0120c; (7) PFA0125c; (8) PFA0130c. Our results show that the energy profile correlates well with our experimental nucleosome landscape (average correlation coefficient = 0.65, measured using a 10-kb-long nonoverlapping sliding window approach), indicating that DNA bending properties dictates, at least partially, the parasite nucleosome positioning.











