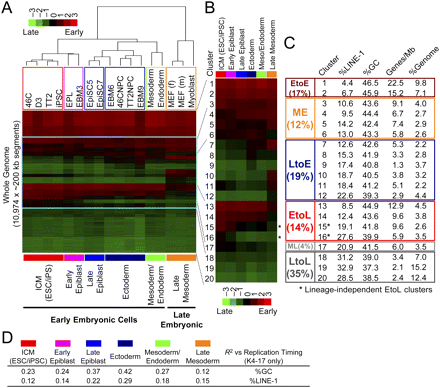
Replication-timing changes across different lineages collectively affect nearly half the genome and include a set of lineage-independent early-to-late changes. (A) Hierarchical clustering with the addition of Gsc+Sox17– mesoderm, Gsc+Sox17+ endoderm, embryonic fibroblasts (f, female; m, male), and fetal myoblast cells. Their characterization is provided in Supplemental Figures 11 and 12. K-means clustering was applied first (K = 20; these K-means clusters are different from those defined in Fig. 5C) and for hierarchical clustering, K-means clusters 4–17 (framed in blue) were used, which showed a differential of >0.80 between any cell types. (B) Centroids (a set of average replication-timing ratios for a given cluster) of K-means clusters 1–20 in A presented in a heatmap format, which shows the average ratios for six cell states. Asterisks indicate lineage-independent EtoL clusters. (C) Sequence properties of K-means clusters 1–20 in A and B. Based on the average replication-timing ratios, we categorized clusters into those that stay early (EtoE), late (LtoL), middle-early (ME), middle-late (ML), shift earlier (LtoE), and shift later (EtoL). LtoE and EtoL clusters were defined using a stringent cutoff (those with a differential of >1.0). (D) Correlation between replication timing and GC/LINE-1 content in different cell states. Clusters 4–17 were used for calculation of Pearson's R2-values.











