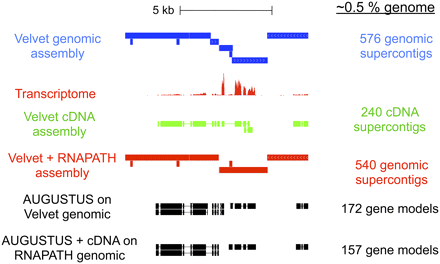
An example of Velvet supercontigs (blue) and RNAPATH supercontigs (red) on 20 kb of Sanger-sequenced PS1010 fosmids (Kuntz et al. 2008) along with the ERANGE mapped coverage of the transcriptome reads (red), Velvet assembly of transcriptome (green), AUGUSTUS gene predictions on the original Velvet assembly and AUGUSTUS with Velvet-computed cDNA sequences assisted predictions on the final cDNA-scaffolded assembly. cDNA-mediated scaffolding combined with a genefinder improves the accuracy of gene models by allowing genes fragmented between genomic supercontigs to be on the same scaffold (broken line box). Summary statistics on the right are for the entire Sanger-sequenced 429 kb of sequence (corresponding to ∼0.5% of the genome).











