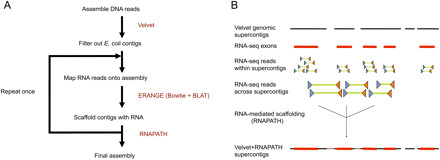
Sequencing strategy. (A) Genomic reads are assembled using the Velvet short read assembler and then filtered for high similarity to E. coli. RNA-seq paired reads are mapped using a combination of Bowtie and BLAT onto this preliminary genomic assembly. The RNA-seq reads are then imported into ERANGE, where those reads with ends on separate supercontigs serve as input to the RNAPATH module. This process can be repeated with trimmed reads to increase mappable reads, if necessary. (B) Paired RNA reads (each pair represented by a blue and an orange triangle connected by a green line), where each read end maps to a separate genomic supercontig, can be used to scaffold (i.e., to join, order, and orient) those genomic supercontigs based on the two read ends pointing toward each other.











