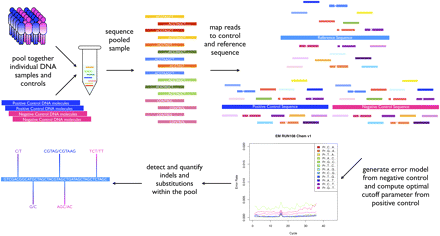
Experimental and computational pipeline for detection of indels and substitutions in large pooled DNA samples: DNA samples from a selected group of patients are individually pooled in a complex mixture to be used as a template for PCR amplification of selected genomic loci. The pool PCR products are then combined in an equimolar mix containing a DNA fragment without variants (negative control) and a synthetic pool with engineered mutations present at the lowest expected variant frequency present in the sample (positive control). The mix is then sequenced on Illumina Genome Analyzer LIX, and sequencing reads are mapped back to the sample and the controls reference sequence by gapped alignment. The negative control reads are used to generate a second-order error model to be used in the variant calling phase. The positive control allows determination of the optimal cutoff for maximizing specificity and sensitivity of the analysis. SPLINTER will then be used to analyze the pooled sample, resulting in detection and quantification of indels and substitutions present in the pool. The SPLINTER algorithm detects true segregated variants by comparing the frequency vector of observed read bases to an expected frequency vector defined by the error model. If the observed vector is significantly different from the expected vector, then SPLINTER will call that position a sequence variant. For each identified variant, SPLINTER will then perform maximum likelihood fit in order to estimate its frequency in the pooled sample.











