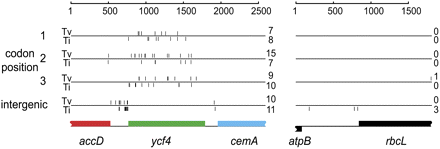
Sequence divergence between L. latifolius and L. cirrhosus in the accD-ycf4-cemA region (left) and the atpB-rbcL region (right). Vertical tickmarks indicate the locations of each nucleotide substitution, categorized according to whether it occurs at codon position 1, 2, or 3; or in intergenic DNA; and as a transversion (Tv; tickmarks above the horizontal lines) or a transition (Ti; tickmarks below the horizontal line). The total numbers of each type of substitution are shown on the right. Supplemental Figure S5 shows the nucleotide sequence alignment summarized in the left panel.











