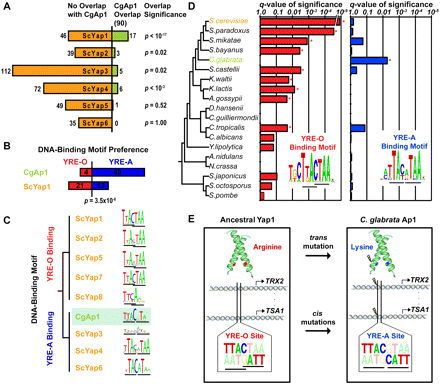
The CgAp1 transcriptional network has been rewired. (A) For the promoters targeted by each yAP-1 transcription factor in Sc, the overlap with CgAp1 targets is shown (of 90 CgAp1 targets total). (B) CgAp1 prefers YRE-A-binding sites compared with ScYap1 (Fisher's exact test). (C) The CgAp1 DNA-binding motif (green) clusters with YRE-A rather than YRE-O motifs. (D) The YRE-O site is enriched (star) among common ScYap1 and CgAp1 targets in other yeasts (hypergeometric test, Q < 0.05), but not Cg (Q ∼ 1.0). The YRE-A site is enriched among these targets in Cg (star) but not other yeasts. (E) Compensatory mutations in both trans and cis maintain AP-1 binding.











