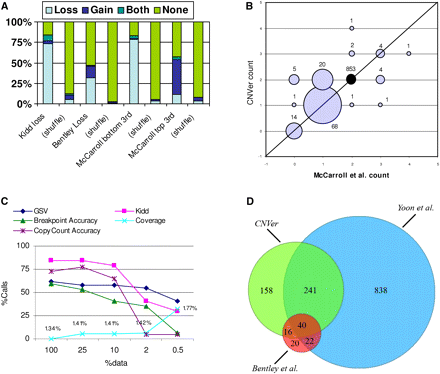
(A) Sensitivity analysis. We compared against four datasets: losses from Kidd et al. (2008) (141 calls) and Bentley et al. (2008) (1933 calls), as well as top and bottom 3rd quantiles from McCarroll et al. (2008) (93 and 26 calls, respectively). The chart shows the percentage of given calls that overlap only CNVer loss calls, only gain calls, both gain and loss calls, and no calls. In the case of Kidd et al., we require that the CNVer call is completely contained within the Kidd et al. call. We also make the same comparison against a randomly shuffled version of our calls. The raw numbers for this chart are included in Supplemental Table 5. (B) Count accuracy. A bubble chart comparing the copy counts reported by McCarroll et al. (2008) with those of CNVer for the 976 regions that do not overlap a segmental duplication. The area of each bubble is proportional to the number of regions with the given joint copy counts (except for the bubble with both counts being 2). One outlier is also not shown, where the CNVer copy count is 10 and the McCarroll et al. count is 2. The diagonal line represents regions where the predictions of the two methods matched. (C) Effect of number of reads. We measure the accuracy of our algorithm on datasets with 100%, 25%, 10%, 2%, and 0.5% of the original mate pairs. We show the percentage of called bases overlapping the GSV, the percentage of Kidd et al.'s calls that we overlap, the percentage of Kidd et al.'s sequenced variants that we detect with an F-score ≥ 0.9 (breakpoint accuracy), the percentage of McCarroll et al.'s regions (out of the 118 with copy count different from 2) for which CNVer's copy count agrees with McCarroll et al. (copy count accuracy), and the percent increase in the number of bases called with respect to the 100% run (coverage). The last series is also marked with the percent of the autosomal genome annotated as copy number variant. (D) Comparison against other methods. A three-way comparison of the calls made by CNVer, Yoon et al. (2009), and Bentley et al. (2008). Two calls are considered to overlap if they share at least one base pair. Note that the overlap measure is not symmetric, e.g., 14 of Yoon et al.'s calls overlap Bentley et al.'s and not CNVer, but 30 of Bentley et al.'s calls overlap Yoon et al.'s and not CNVer. What we show in the intersections, therefore, are the averages.











