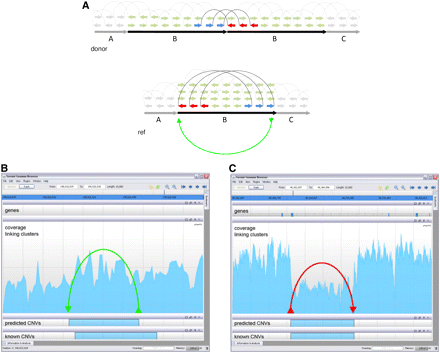
Depth-of-coverage and linking clusters. (A) A tandem duplication changes the DOC and creates discordant mate pairs. The reference genome consists of the segments ABC, while the donor has ABBC. The mate pairs are shown along the donor as they are sequenced and on the reference as they are mapped. The green (light) mate pairs are concordant, with the correct orientation and distance; however, the three bold mate pairs that span the adjacency between the two Bs are discordant. They form a linking cluster that indicates a putative adjacency in the donor from the end of B to the beginning of B (shown by the green edge). In addition, the duplication affects the DOC, as the density of reads mapping to the B segment is twice as high as in the rest of the genome. (B) A screen shot of the Savant (Fiume et al. 2010) genome browser (http://compbio.cs.toronto.edu/savant) on a 9-kb region of chromosome 1, showing the DOC and the linking clusters found by CNVer in our dataset, the gain call CNVer predicts, and an overlapping call from the GSV database of known variants. The underlying variation is likely a tandem duplication, like the one shown in A, because of the clear increase in the DOC and the presence of the green linking cluster. (C) A 19-kb region of chromosome 7 containing a deletion validated by sequencing (Kidd et al. 2008). There is a clear drop off in the DOC, as well as a linking cluster that connects the region left of the deletion to the one right of the deletion. Despite the noise in the DOC signal, the linking cluster allows CNVer to discern the breakpoints of the 10-kb deletion: they are predicted to within 28 bp on the left and 3 bp on the right.











