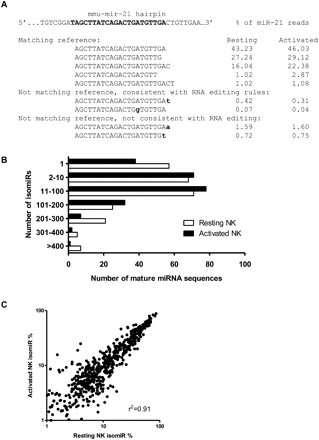
Mature miRNA sequence variation (isomiRs) detected by Illumina GA sequencing of NK cell miRNAs. (A) Example miR-21 isomiRs illustrate the complexity of miRNA sequences identified by Illumina sequencing. Shown are selected isomiR sequences that match or do not match the reference genome and their percent contribution to miR-21 read counts (mature miR-21 sequence annotated in bold). We further divide those reads not matching the reference genome into sequences that may arise by a known RNA editing event and all others. A complete breakdown of isomiR sequence variation for all miRNAs is provided in the Supplemental material. (B) Distribution of miRNA mature sequences by the number of isomiRs identified by Illumina GA sequencing. (C) Comparison of isomiR sequences between resting and activated NK cell libraries. Scatterplot showing the percent contribution of a given isomiR in each library, with each dot representing one isomiR. Analysis was limited to the top five isomiRs and the top 125 miRNAs by expression. In general, isomiRs were found in very similar distributions in resting and activated NK cells (r2 = 0.91).











