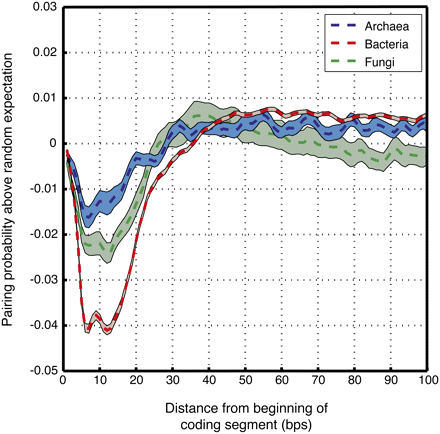
Coding regions tend to encode depletion of RNA secondary structure downstream of the start codon. Shown is the difference in the probability of being base-paired between the real and randomized genomes, averaged across the first 100 nt of all coding segments of archaeal (blue), bacterial (red), and fungal (green) genomes. The patches show SE of the difference. Pairing probabilities were predicted by using the Vienna package (Hofacker 2003) to fold the real and randomized genomes. Each curve was smoothed with a 3-bp moving window. Since the first codon in all coding segments has only one flanking codon, it is never swapped by our genome randomization method. Thus, by construction, the first nucleotides of the coding region are more similar between the real and randomized genomes, explaining the lower difference observed in the pairing probability of these nucleotides between the real and randomized genomes.











