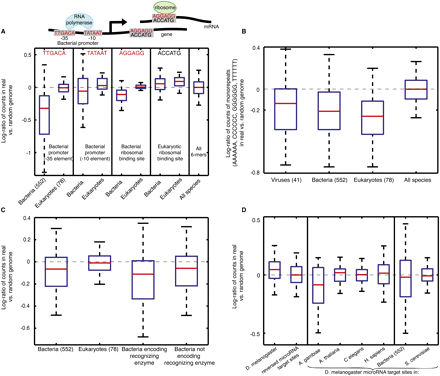
Coding sequences display phyla-specific enrichment of known biological signals. (A) Box plot of log-ratio of number of appearances between real and randomized genomes, of sequence determinants of transcription (−35 promoter element, TTGACA; −10 promoter element, TATAAT) and translation initiation in bacteria (Shine-Dalgarno motif, AGGAGG) and of translation initiation in eukaryotes (Kozak motif, ACCATG). In each species, each of the above 6-mer sequences is counted out-of-frame in the real genome and in the randomized genome, and the log-ratio of these counts is incorporated into the box plot. The red line denotes the median, the blue box delimits 25–75 percentiles, and the outermost bars show the minimum and maximum. The number of species from each phyla group is shown in parentheses. (B) Same as A, for log-ratios of mononucleotide 6-mers across various phyla. All represents all n-mers in all species. (C) Same as A, for bacterial restriction enzyme sites. The bacteria encoding recognizing enzyme group (third box plot from left) only displays log-ratios of restriction enzyme sites in bacterial genomes that encode the enzymes that recognize those sites, whereas the bacteria not encoding recognizing enzyme group (rightmost box plot) only displays log-ratios of restriction sites in bacterial genomes that do not encode the recognizing enzymes. (D) Same as A, for log-ratios of microRNA target sites from Drosophila melanogaster. The 7-mer seed (reverse complement of nucleotides 2–8 of the microRNA) from each microRNA was taken for the log-ratio computation. The log-ratios are shown in the coding sequences of Drosophila and in several other species, as well as in 552 bacterial genomes. The distribution of the reverse sequences of the microRNA target sites is also shown as a control.











