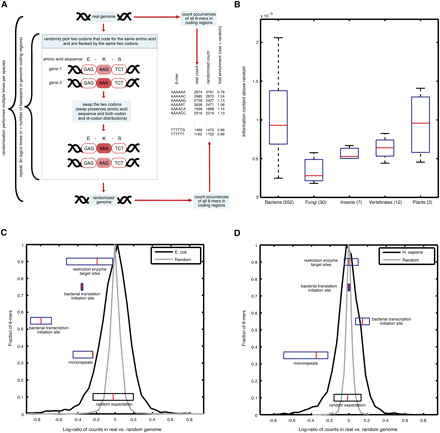
Overview of our approach and detection of additional information encoded within protein-coding sequences. (A) Illustration of our method for identifying over- and underrepresented short sequences within coding regions. For each short sequence (here 6-mer sequences are shown), we count the number of its appearances in a given genome's coding sequences and compare that to its average number of appearances in the coding sequences of randomized genomes. The randomization swaps codons from different genomic locations only if they are both flanked by identical codons and, thus, preserves amino acid sequence, codon usage, and di-codon counts. An example of one codon swap is shown (left), and these swaps are repeated iteratively for each randomization, for each species. (B) All genomes contain additional information in their coding sequences. Shown is the Jensen-Shannon information divergence, a measure analogous to information content, between the distribution of all 6-mer sequences when counted out-of-frame in the real and randomized genomes (since our randomization preserves di-codon counts, the counts of 6-mers in-frame are equal in the real and random genomes, by construction). The Jensen-Shannon divergence is shown as a box plot for all organisms in various phyla groups. The red line denotes the median, the blue box delimits 25–75 percentiles, and the outermost bars show the minimum and maximum. The number of species from each phyla group is shown in parentheses. (C) Histograms of log-ratios of number of appearances of the out-of-frame 6-mers in E. coli (black) and out-of-frame 6-mers in randomized E. coli genomes (gray). Box plots of log-ratios for specific families of known biological signals (mononucleotide repeats, restriction enzyme target sites, and bacterial transcription and translation initiation sites) are shown in their appropriate place along the histogram. Histograms were normalized to have a maximum of 1 for ease of comparison. (D) Same as C, but for human.











