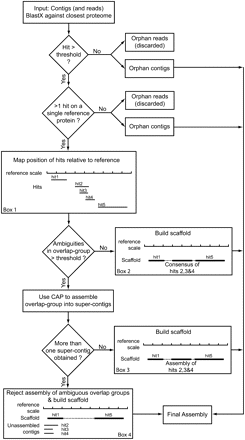
Diagrammatic representation of the STM method. This pipeline can either use only contigs (STM- method) or, if reads are long enough, contigs plus unassembled reads (STM+ method). These contigs/reads are mapped on the reference proteome using BLASTX. When a contig has no significant hit or is the only one to map on a given reference protein, it cannot be further assembled and is directed into the final assembly. When there are several hits on a same reference protein (Box 1: an example with 5 hits) their relative positions are recorded on the reference scale. If there is an overlap in the positioning of several hits (here hits 2, 3, and 4 form an overlap group), their consensus sequence is computed, and when the number of ambiguities is below a user-defined threshold, the consensus is accepted and a scaffold is constructed (Box 2: dashed line represents N's added to join the contigs). Else, the consensus is rejected and the contigs of the overlap group are assembled using CAP. If the result of this assembly step is a single “super-contig,” it is accepted and a scaffold is constructed (Box 3). If more than one super-contig is obtained (Box 4), the overlap group assembly is rejected and the contigs are placed as independent transcripts in the final assembly. If present, the other nonoverlapping hits (or nonambiguous overlap groups) are joined into a scaffold, which is incorporated into the final assembly.











