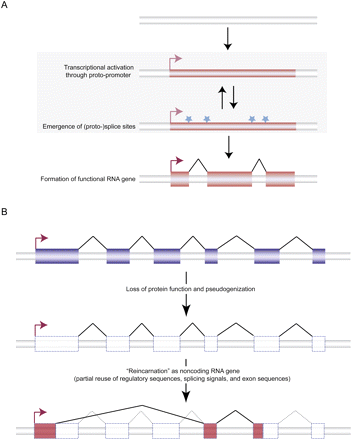
Evolutionary origins of long noncoding RNA genes. (A) De novo emergence. In this scenario, previously nonfunctional genomic sequence becomes transcribed (thin red box) through the acquisition/activation of a proto-promoter sequence (right-angled arrows). The transcriptional activation may be followed or preceded by the evolution of (proto-) splice sites (light blue stars). Together, these events allow for the formation of potentially functional and selectively beneficial multi-exonic noncoding RNA genes. (Large red boxes) Exons, (thin black lines) splicing, (red right-angled arrows) TSSs. (B) Origin of noncoding RNA gene from ancestral protein-coding gene. In this process, the original (functionally redundant) protein-coding gene loses its function and becomes a pseudogene. After or during loss of protein function and coding exon decay, a new functional noncoding RNA gene may arise, a process that may draw from regulatory elements and other sequences (splicing signals, exon sequences, polyadenylation sequences, etc.) from the ancestral protein-coding gene. (Blue boxes) Protein-coding exons, (red boxes) RNA exons, (transparent boxes) pseudogenized exons, (thin black lines) splicing, (dotted lines) lost ancestral splicing capacity, (red right-angled arrows) TSSs.











