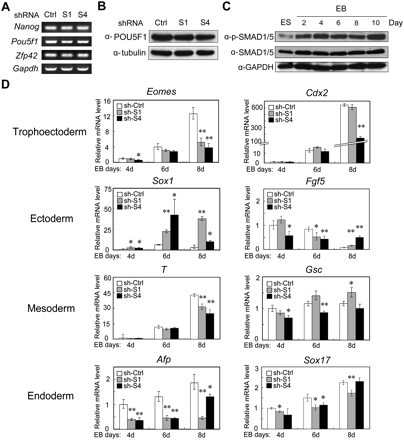
SMAD1 and SMAD4 are largely involved in differentiation regulation rather than direct self-renewal maintenance. (A) Knockdown of Smad1 or Smad4 has no effect on the expression of self-renewal markers. Total RNA extracted from R1 cells expressing Smad shRNA or control shRNA constructs were subjected to RT-PCR. Gapdh served as a loading control. (B) No difference of the POU5F1 protein levels in R1 cells expressing Smad shRNA or control shRNA. Protein expression was examined by immunoblotting. Tubulin served as a loading control. (C) Phosphorylation of SMAD1/5 is enhanced during EB formation. EB was formed in serum-free KO-SR culture conditions, and total proteins were collected at indicated times and subjected for anti-phopho-SMAD1/5 and anti-SMAD1/5 immunoblotting. GAPDH served as a loading control. (D) Smad knockdown alters the expression profile of germ layer markers. Quantitative RT-PCR analysis was performed to examine marker expression in Smad shRNA and control shRNA cells during the course of EB differentiation. The significance of expression was analyzed by Student's t-test, and data are presented as mean ± SEM (n = 3; **P < 0.01; *P < 0.05). This experiment was repeated three times and similar results were obtained.











