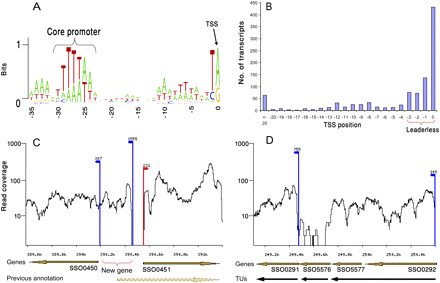
The structure of the S. solfataricus transcriptome. (A) Core promoter. Positions relative to TSS are marked below the sequences. The height of each letter corresponds to its frequency at that position. The core promoter motif in Archaea is indicated. The plot was prepared using the WebLogo software tool (Crooks et al. 2004). (B) Distribution of mapped TSS positions relative to the ORF ATG codon (n = 1040). Position 0 depicts transcripts beginning exactly at the adenine of the ATG. (C) Example for correction of gene annotations. (Dashed arrow) Marks the previous annotation of gene SSO0450 (conserved protein implicated in secretion) that was computationally predicted to begin in position 390,329. Transcriptome data indicate that it actually begins 228 bp downstream. A new protein-coding gene, having homology with a small heat-shock protein Hsp20 in other archaea, is identified on the reverse strand. Read coverage is in log scale. (Red arrows) TSS on the forward strands; (blue arrows) TSS on the reverse strands. The number above TSS indicates the number of reads supporting transcript beginning at that position. (D) Definition of transcriptional units (TUs). Operon annotation is refined based on continuous expression and TSS existence. Four closely spaced genes might be predicted to appear in the same operon based on the distance between them. Transcriptome data show at least two separate TUs. A third TU, corresponding to gene SSO5576, does not show expression and might either represent an unexpressed gene or a spurious gene prediction.











