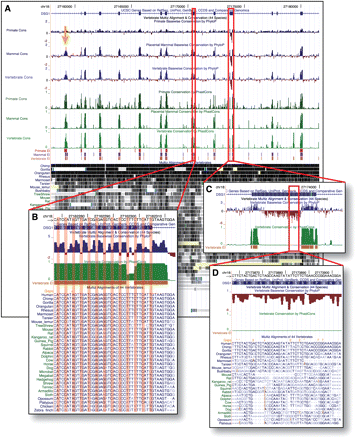
Conservation track in UCSC Genome Browser. A portion of the desmoglein 1 (DSG1) gene on human chromosome 18 shown with the new Conservation track, including a 44-way vertebrate alignment and nine conservation subtracks. The subtracks display phyloP scores (in blue and red), phastCons scores (green), and phastCons-predicted conserved elements (pink, purple, and mustard) for all species, the 32 placental mammals, and the nine primates (bottom to top within each group). (A) The phyloP and phastCons scores are broadly similar when the display is zoomed out, with scores near zero for most noncoding regions but elevated in exons (thick blue bars at top) as well as in conserved noncoding elements (orange arrow). (B) At finer resolution, however, phyloP reveals significantly more variation from base to base than does the hidden Markov model–based phastCons. In this coding exon, codon position effects are clearly evident from phyloP but not from phastCons. (C,D) The phyloP tracks also indicate accelerated evolution (with negative scores, shown in red), while phastCons measures conservation only. Here an exon with a striking fast-evolving segment is shown. Interestingly, cDNA data from other mammals suggest that this exon derives from a fusion of two ancestral exons, with the fast-evolving segment corresponding to the ancestral intron.











