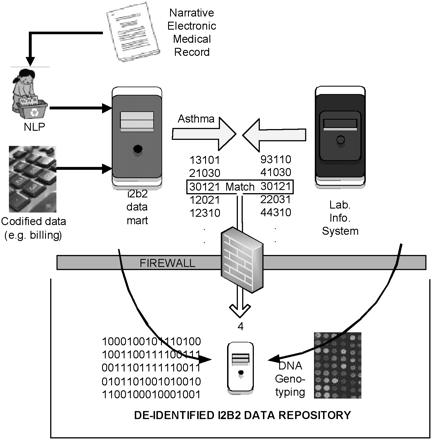
Matching anonymously identified populations to anonymous samples. An i2b2 datamart is generated from codified data (e.g., billing codes, laboratory test values) and concepts codified by running the narrative text in electronic medical records through a NLP tool, the HITEx package described in the Methods section. Patients included within the i2b2 datamart meeting study criteria are selected and their corresponding set of identifiers are generated. Those identifiers are forwarded to the Crimson application, which scans recent transactions forwarded from one or more local clinical laboratory or pathology information systems to identify newly accessioned materials matching the cohort identifiers and desired sample types. Upon completion of diagnostic testing (1–3 d after collection in most cases), Crimson manages reaccession of the sample to a study's IRB protocol and assigns the i2b2-forwarded subject ID to a uniquely generated sample ID. These actions remove all identifiers (accession no., medical record no., etc.) from the original sample. The sample may then be released to the investigator where it can be measured (for genome-wide genotyping in this instance), and these measurement data are merged with the phenotypic data set in the i2b2 datamart. Because of the electronic and regulatory firewalls, only research personnel approved by the IRB can view the limited data set (in the HIPAA sense), and they cannot view the identified clinical data visible to those who access the laboratory information system.











