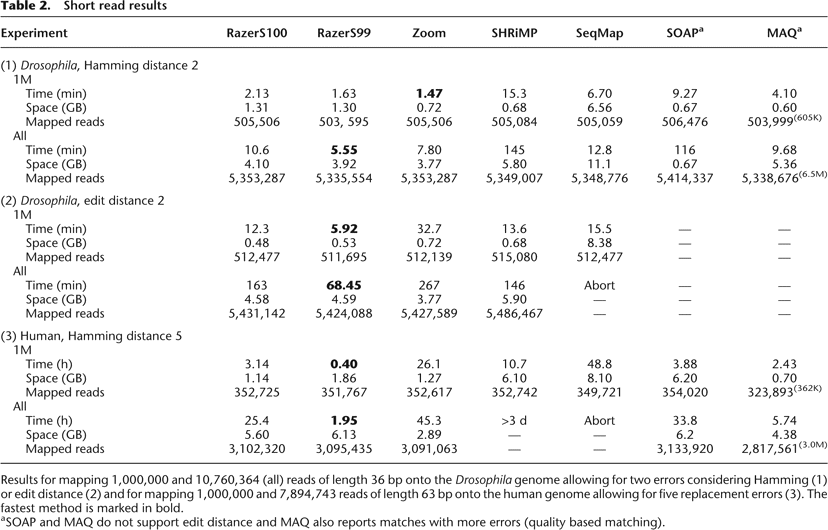
Short read results
Click on table to view larger version.
-
Results for mapping 1,000,000 and 10,760,364 (all) reads of length 36 bp onto the Drosophila genome allowing for two errors considering Hamming (1) or edit distance (2) and for mapping 1,000,000 and 7,894,743 reads of length 63 bp onto the human genome allowing for five replacement errors (3). The fastest method is marked in bold.
-
aSOAP and MAQ do not support edit distance and MAQ also reports matches with more errors (quality based matching).











