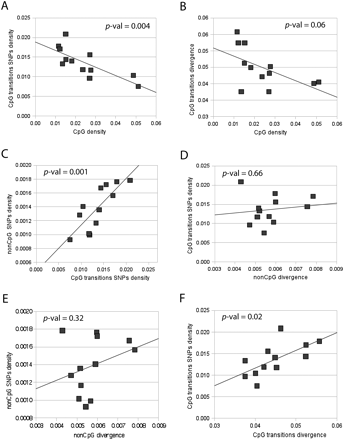
Correlations between polymorphism, interspecies divergence, and CpG content. We analyzed divergence in the chimpanzee lineage after divergence from human using orangutan as an outgroup in order to compensate for bias due to padlock probe design based on the presence of CpGs in the human sequence. SNP densities were calculated as normalized densities per site of a specific type. To calculate the density of CpG SNPs, we divided the total number of the observed CpG polymorphisms in the region by the combined length of all surveyed CpG nucleotides in the region. Correspondingly, to calculate the density of non-CpG SNPs, we divided the total number of observed non-CpG polymorphisms in the region by the combined length of all surveyed non-CpG nucleotides in the region. (A) Correlation between densities of SNPs originated as transitions in CpG sites and fraction of CpGs in the region. (B) Correlation between substitutions due to CpG transitions in the chimpanzee lineage after divergence with humans and fraction of CpGs in the region in the human genome. (C) Correlation between densities of SNPs originated as transitions in CpG sites and non-CpG SNP density. (D) Correlation between densities of SNPs originated as transitions in CpG sites with non-CpG divergence in the chimpanzee lineage after split with humans. (E) Correlation between non-CpG SNP density with non-CpG divergence in the chimpanzee lineage after split with humans. (F) Correlation between densities of SNPs originated as transitions in CpG sites with divergence in the chimpanzee lineage due to CpG transitions.











