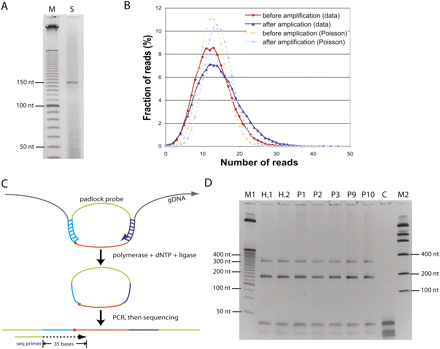
Padlock probe capture of 53,777 CpG sites. (A) The raw probe precursor (150-mer) sample from Agilent (S) was loaded along with a 10-bp ladder (M) on a 6% denaturing PAGE gel. (B) The probe precursors before and after two rounds of PCR amplification were end-sequenced by Illumina Genome Analyzer. (C) The padlock probes were hybridized to the targeted genomic CpG sites with a uniform 40-nt size. To simplify library construction, a target CpG (dot) was located immediately next to the ligation arm of the probe. Enzymatic filling and ligation of the gap (brown) allowed a copy of the target site to form a circle with the padlock probe. The circles were then PCR amplified using the backbone sequences (green) as primers. The common backbone sequence immediately upstream of the ligation arm served as a sequencing primer. (D) Amplification of circles derived from padlock probes. PCR products were loaded on a 6% PAGE gel. The two upper DNA bands had the expected amplicon sizes: 184 bp (subject to gel purification and Illumina sequencing) and 334 bp (if polymerization extended around the circle twice); the lower bands below 50 nt were derived from PCR primers. (Lane M1) 25-bp DNA ladder (Invitrogen); (lanes H.1,H.2) technical replicates of HapMap sample NA10835; (lanes P1,P2,P3,P9,P10) Personal Genomes 1, 2, 3, 9, and 10, respectively; (lane C) no genomic DNA control; (lane M2) low mass DNA ladder (Invitrogen).











