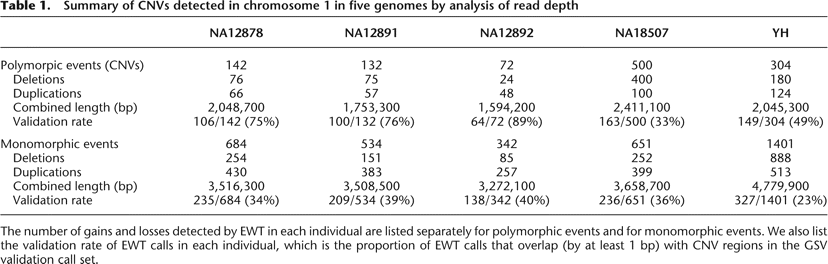
Table 1.
Summary of CNVs detected in chromosome 1 in five genomes by analysis of read depth
Click on table to view larger version.
-
The number of gains and losses detected by EWT in each individual are listed separately for polymorphic events and for monomorphic events. We also list the validation rate of EWT calls in each individual, which is the proportion of EWT calls that overlap (by at least 1 bp) with CNV regions in the GSV validation call set.











