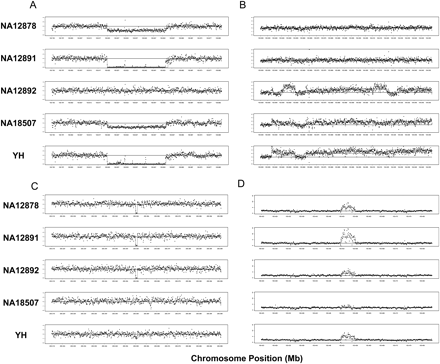
Examples of CNVs detected by analysis of RD. We present four examples of polymorphic gains and losses detected by EWT in five individuals. The x-axis represents genomic coordinates (in Mbp) and the y-axis represents RD, which is median-normalized to copy number 2. In each panel, plots are for NA12878, NA12891, NA12892, NA18507, and YH from top to bottom. The coordinates of A, B, C, and D are chr1:150,792,101–150,884,101, chr1:103,930,401–104,053,201, chr1:205,319,001–205,399,701, and chr1:150,422,701–150,486,501, respectively.











