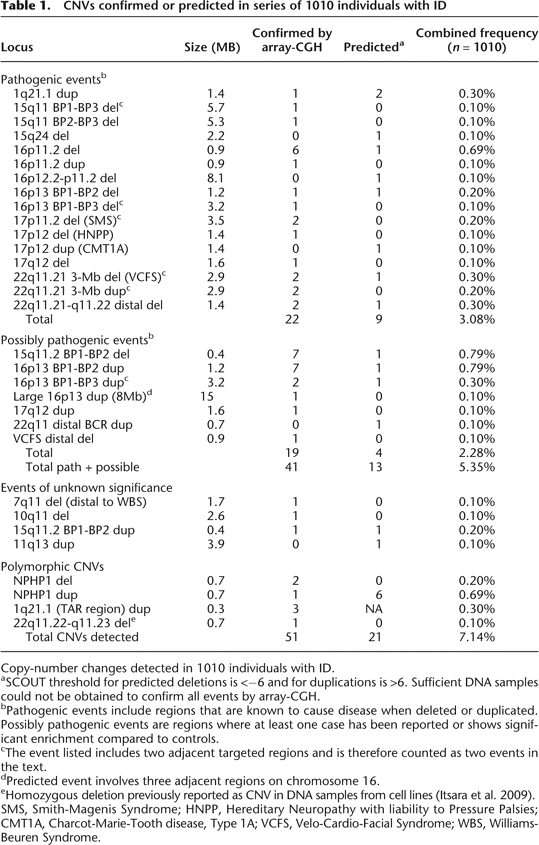
CNVs confirmed or predicted in series of 1010 individuals with ID
Click on table to view larger version.
-
Copy-number changes detected in 1010 individuals with ID.
-
aSCOUT threshold for predicted deletions is <−6 and for duplications is >6. Sufficient DNA samples could not be obtained to confirm all events by array-CGH.
-
bPathogenic events include regions that are known to cause disease when deleted or duplicated. Possibly pathogenic events are regions where at least one case has been reported or shows significant enrichment compared to controls.
-
cThe event listed includes two adjacent targeted regions and is therefore counted as two events in the text.
-
dPredicted event involves three adjacent regions on chromosome 16.
-
eHomozygous deletion previously reported as CNV in DNA samples from cell lines (Itsara et al. 2009).
-
SMS, Smith-Magenis Syndrome; HNPP, Hereditary Neuropathy with liability to Pressure Palsies; CMT1A, Charcot-Marie-Tooth disease, Type 1A; VCFS, Velo-Cardio-Facial Syndrome; WBS, Williams-Beuren Syndrome.











