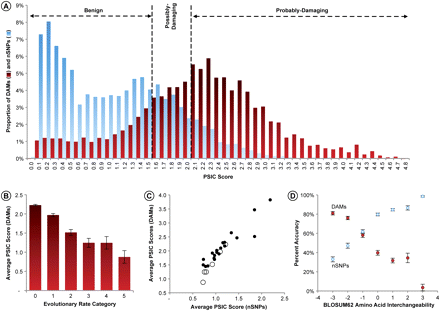
(A) Frequency distributions of DAM (red) and nSNPs (blue) PSIC scores. Vertical lines show the PolyPhen PSIC cut-offs for classification of variants in the absence of structural or other information; nSNPs and DAMs are from Subramanian and Kumar (2006a). (B) Mean PSIC values for DAMs in different evolutionary rate categories. The correlation (r) between mean PSIC values and mean evolutionary rate is −0.96 (P << 0.01). The 95% confidence intervals derived from the SEMs are shown. (C) Relation between mean PSIC scores for DAMs and mean PSIC scores for nSNPs, by amino acid types (solid circles) and evolutionary rates (open circles). (D) Inverse relationship of the accuracy of DAMs (probably-damaging) and nSNPs (benign) with the evolutionary interchangeability of amino acid pairs (original/variant pairs) as captured in the BLOSUM62 matrix. Each data point represents the average of all pairs for a given BLOSUM score, with the error bars displaying the 95% confidence intervals derived from binomial variance of the proportions. BLOSUM scores are log-odds substitution occurrences. Negative BLOSUM scores show amino acid pairs that are found to have a low probability of substitution, whereas a positive score indicates frequently observed amino acid pairs. Complete 20 × 20 matrices of DAM and nSNP accuracies (and their SEs) are given in the Supplemental material.











