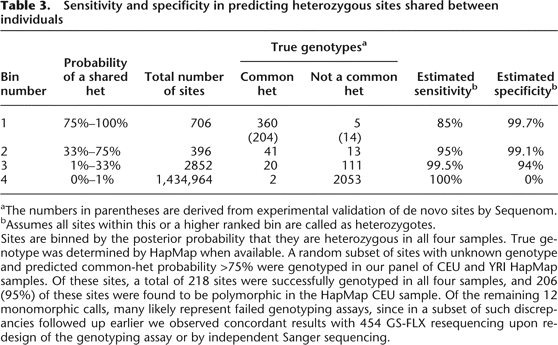
Sensitivity and specificity in predicting heterozygous sites shared between individuals
Click on table to view larger version.
-
aThe numbers in parentheses are derived from experimental validation of de novo sites by Sequenom.
-
bAssumes all sites within this or a higher ranked bin are called as heterozygotes.
-
Sites are binned by the posterior probability that they are heterozygous in all four samples. True genotype was determined by HapMap when available. A random subset of sites with unknown genotype and predicted common-het probability >75% were genotyped in our panel of CEU and YRI HapMap samples. Of these sites, a total of 218 sites were successfully genotyped in all four samples, and 206 (95%) of these sites were found to be polymorphic in the HapMap CEU sample. Of the remaining 12 monomorphic calls, many likely represent failed genotyping assays, since in a subset of such discrepancies followed up earlier we observed concordant results with 454 GS-FLX resequencing upon redesign of the genotyping assay or by independent Sanger sequencing.











