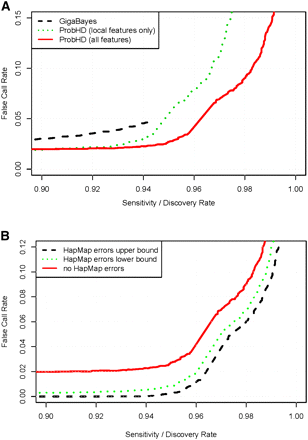
(A) Ability of three classifiers to identify known heterozygous sites. False call rate (FCR) (the fraction of called heterozygous SNPs that are known to be homozygous) is shown as a function of sensitivity (the fraction of known heterozygous sites called by each classifier). Three classifiers are compared: (1) GigaBayes; (2) ProbHD local-feature classifier, which considers all local features that could be extracted from the 454 GS-FLX generated MSA and quality scores file; and (3) ProbHD full classifier, which considers both local- and amplicon-level features from alignments generated by hAlign. (B) Estimated sensitivity and FCR for calling a site heterozygous, corrected for HapMap errors. (Dashed line) Assumes a HapMap error rate at the upper end of the 95% confidence interval; (dotted line) lower end; (solid line) no HapMap errors.











