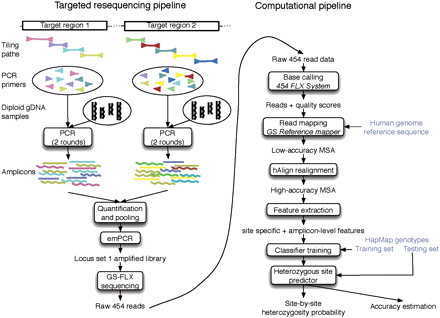
Graphical overview of the four steps in the prediction pipeline. (1) Sequencing: Target regions are amplified by LR-PCR; amplicons are sequenced using a 454 GS-FLX sequencer. A set of sequence reads is generated by the 454 GS-FLX base-caller. (2) Alignment: Reads are aligned to the reference sequence and combined into a multiple sequence alignment (MSA). (3) Feature extraction: Numerical features are computed from the MSA for each site in the target region. (4) Training: Given a training set of sites with known genotypes from the HapMap database, we train a classifier to identify heterozygous sites from sequencing data. This classifier is then applied to novel data sets to identify novel SNPs.











