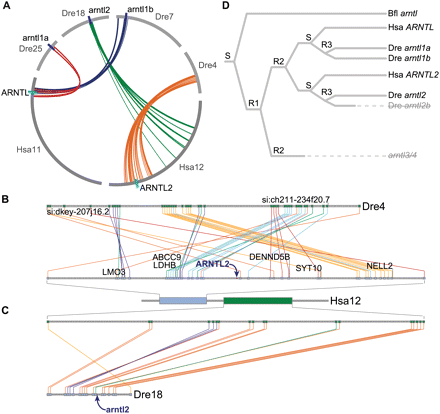
Conserved syntenies for ARNTL genes. (A) A circle plot summarizing human and zebrafish ARNTL family clusters. Arcs along the circumference of the circle represent chromosomes, while arcs within the circle connect pairs of orthologs. (B) The ARNTL2 orthologous syntenic cluster showing strong syntenic conservation between Hsa12 and Dre4. Several genes that are part of the original Hsa11/Hsa12 paralogous cluster (Fig. 2C) are labeled. A transposition moved two parts of the Dre4/Hsa12 cluster relative to one another (orange and blue lines). The fourth ARNTL gene in zebrafish (putative arntl2b) would have existed directly upstream of either si:dkey-207j16.2 or si:ch211-234f20.7 on Dre4 before its loss. (C) The arntl2 orthologous syntenic cluster showing syntenic conservation between portions of Hsa12 and Dre18. The zebrafish arntl2 gene did not appear in this cluster because the pipeline misidentified it (see text); its position in the cluster is marked with an arrow. Human orthologs in the Dre18/Hsa12 cluster fall ∼25 Mb from ARNTL2 on Hsa12 (Fig. 2D) due to an inversion occurring after the zebrafish and human lineages diverged. (D) A gene tree showing the inferred evolutionary history of the ARNTL gene family in the amphioxus (Bfl), zebrafish (Dre), and human (Hsa) lineages. (S) A speciation event; (R1, R2, R3) three whole-genome duplications in the lineages leading to human and zebrafish. Genes in pale, strikethrough text have been lost.











