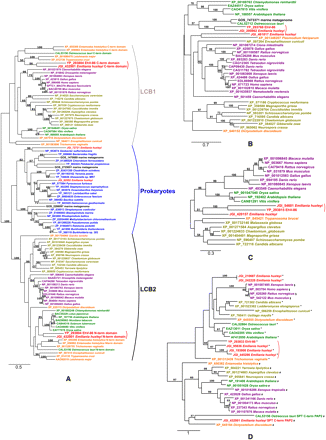
Maximum likelihood phylogenetic trees based on the amino acid sequences of the four central enzymes in the sphingolipid biosynthesis pathway. (A) Serine palmitoyltransferase LCB1 and LCB2 domain sequences and their homologs. (B) Dihydroceramide synthases (LAG1). (C) Dsd1-like fatty acid desaturases. (D) Lipid phosphate phosphatase (LPP), the PAP2-domain sequence from the E. huxleyi SPT, and their homologs. These trees are unrooted per se, although we have arbitrarily chosen a root (mostly by mid-point rooting) for each tree only for visualization purpose. The number of substitutions per site is indicated under the scale bar. In D, (#) sequences best hitting to sphingosine 1-phosphate phosphatases (cd03388) after NCBI/CDD searches; (*) sequences best hitting to phosphatidic acid phosphatases (cd03390); (§) sequences best hitting to the wunen subfamily sequences (a family of membrane associated phosphatidic acid phosphatases; cd03384).











