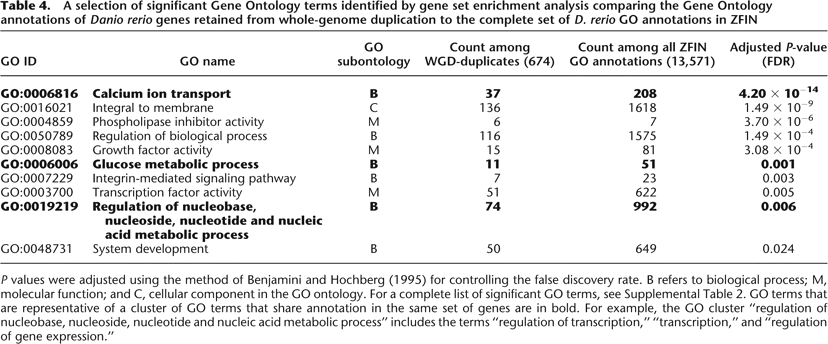
A selection of significant Gene Ontology terms identified by gene set enrichment analysis comparing the Gene Ontology annotations of Danio rerio genes retained from whole-genome duplication to the complete set of D. rerio GO annotations in ZFIN
Click on table to view larger version.
-
P values were adjusted using the method of Benjamini and Hochberg (1995) for controlling the false discovery rate. B refers to biological process; M, molecular function; and C, cellular component in the GO ontology. For a complete list of significant GO terms, see Supplemental Table 2. GO terms that are representative of a cluster of GO terms that share annotation in the same set of genes are in bold. For example, the GO cluster “regulation of nucleobase, nucleoside, nucleotide and nucleic acid metabolic process” includes the terms “regulation of transcription,” “transcription,” and “regulation of gene expression.”











