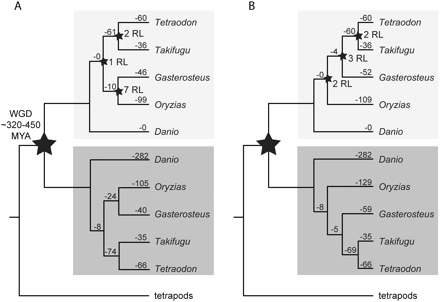
Inferred gene losses along different teleost lineages. For each of the 754 protein families containing a WGD-duplicate, one of the D. rerio duplicate gene copies was arbitrarily designated the reference point against which the presence of the locus in the other fish taxa and in the sister clade was assessed. We marked all instances where a fish taxon or clade was inferred to have lost a copy of the locus, taking into account two plausible species tree topologies: (A) the tree topology as represented in the NCBI taxonomy, which supports a monophyletic clade Smegmamorpha containing Oryzias latipes and Gasterosteus aculeatus, or (B) the tree topology supported by mitogenomic analyses, which resolves O. latipes as immediately basal to a clade containing G. aculeatus and the Tetraodontiformes. Instances where two teleost species had lost alternative copies of the same locus were counted as “reciprocal gene losses” (data not shown). Reciprocal gene losses that are consistent with the loss having occurred at the time of species divergence are marked RL.











