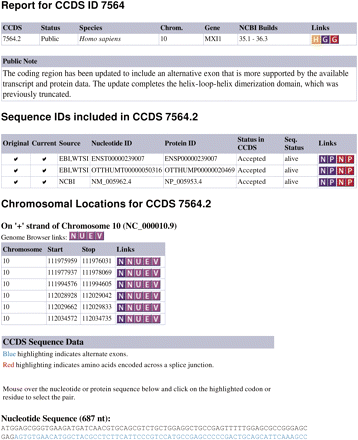
Detailed CCDS ID report page for a MXI1 protein. The CCDS report page presents three tables of information followed by nucleotide and protein sequences for the annotated CDS. The first table summarizes the status for the specified CCDS ID. Colored icons provide links to related resources, to a history report (orange H icon), or to re-query the CCDS database with a different type of identifier. (Red G icon re-queries the CCDS database by GeneID to return all CCDS IDs available for a gene.) A Public Note is provided for a subset of curated records to explain the nature of, and/or the reason for, an update or withdrawal. The sequence identifiers table reports sequences tracked as members of the CCDS ID. A checkmark in the first column (Original) identifies sequence identifiers represented on the annotated genome and included in the analyzed input data sets, and a checkmark in the second column (Current) identifies those that are considered current members (see Supplemental Fig. 2) The Chromosome Locations table reports the genomic coordinates of each exon of the CDS with links to view the annotation in different browsers—the violet icons (N) NCBI; (U) UCSC; (E) Ensembl; and (V) Vega. The nucleotide sequence of the CDS, derived from the genome sequence using the reported exon.











