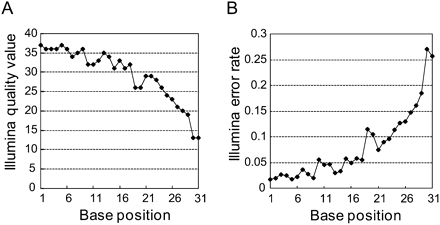
Figure 2.
Distribution of base quality QVs given by Illumina of small RNA reads after trimming of 3′ linker sequences. (A) Average of given Illumina QVs at each base position. (B) Expected Illumina error rate P at each base. P is calculated from Q (the Illumina QV) at each position according to the formula: Perror = 1/[1 + 10(Q/10)].











