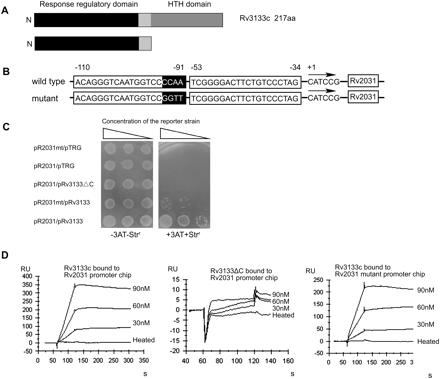
DNA-binding specificity examined by the bacterial one-hybrid reporter system. (A) A 143 amino acid mutant protein of Rv3133c (DevR) was produced by deleting its C-terminal domain and used for the DNA-binding specificity assay. (B) A mutant promoter of Rv2031 was constructed by mutating its CCAA as GGTT, as shown in black background. (C) The bacterial one-hybrid assays. The promoters were cloned into pBXcmT. Rv3133c and its C-terminal deleted mutant, Rv3133ΔC, were cloned into pTRG vectors, respectively. The E. coli XL1-Blue MRF′ Kan strain was used for the routine propagation of all pBXcmT and pTRG recombinant plasmids. A pair of pBXcmT/pTRG plasmids was co-transformed into the reporter strain and then its growth was tested together with the self-activation controls on a selective medium containing 3-AT, Kanr, Strr, and Chlr. Positive growth cotransformants were selected on the selective screening medium plate. (D) SPR assays for protein-DNA interactions. Both Rv3133c and Rv3133ΔC were expressed in E. coli and purified. Biotin-labeled promoter DNA was immobilized on the SA chips (BIAcore). Following a period of stabilization, Rv3133c or Rv3133ΔC protein was passed over the chip. Each analysis was performed in triplicate. An overlay plot was produced for depicting the interaction between Rv3133c or Rv3133ΔC with different concentrations of promoter DNA. RU, response units; s, seconds.











