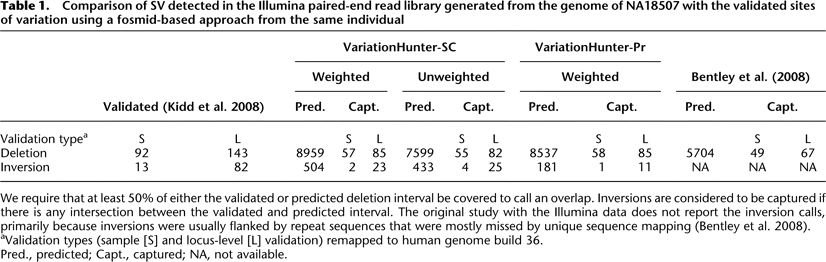
Comparison of SV detected in the Illumina paired-end read library generated from the genome of NA18507 with the validated sites of variation using a fosmid-based approach from the same individual
Click on table to view larger version.
-
We require that at least 50% of either the validated or predicted deletion interval be covered to call an overlap. Inversions are considered to be captured if there is any intersection between the validated and predicted interval. The original study with the Illumina data does not report the inversion calls, primarily because inversions were usually flanked by repeat sequences that were mostly missed by unique sequence mapping (Bentley et al. 2008).
-
aValidation types (sample [S] and locus-level [L] validation) remapped to human genome build 36.
-
Pred., predicted; Capt., captured; NA, not available.











